Epigenetic Regulation of Gene Expression
Epigenetic modifications are added and removed by special classes of enzymes, broadly termed “epigenetic regulators.” Despite extensive study, the full impact of many epigenetic modifications on gene expression is not completely understood, nor is the mechanism of action of epigenetic regulator enzymes which add and subtract the modifications. Our lab seeks to better understand (i) how epigenetic regulators achieve target gene specificity, (ii) mechanisms of epigenetic regulator cellular function and enzymatic activity, and (iii) how the presence or absence of specific epigenetic modifications impacts gene expression, both under standard conditions and under conditions of stress, aging, and disease. We are particularly interested in the mechanisms by which two epigenetic modifier enzymes, Sirt6 and Jim, regulate the epigenome and gene expression in different conditions.
Within the nuclei of our cells, DNA is tightly wrapped and condensed around bead-like structures called nucleosomes, which are made of histone proteins. Histone proteins are modified by a variety of chemical marks, including acetylation and methylation, that together form the “histone code,” a type of epigenetic regulatory system that helps control DNA structure and gene expression.
These modifications are added and removed by specialized enzymes known as writers and erasers, and are interpreted by reader proteins that recognize specific marks and recruit regulatory complexes. Through this coordinated system, histone modifications promote either gene activation or repression.
Among these modifications, histone acetylation is generally associated with active gene expression. Acetylation levels are dynamically regulated by the opposing actions of histone acetyltransferases and histone deacetylases. Sirt6 is a highly conserved (present in multiple species) histone deacetylase, which removes acetyl groups from specific histone residues to fine-tune gene expression programs. In addition, Sirt6 exhibits mono-ADP-ribosylase activity, making it a multifunctional epigenetic regulator enzyme.
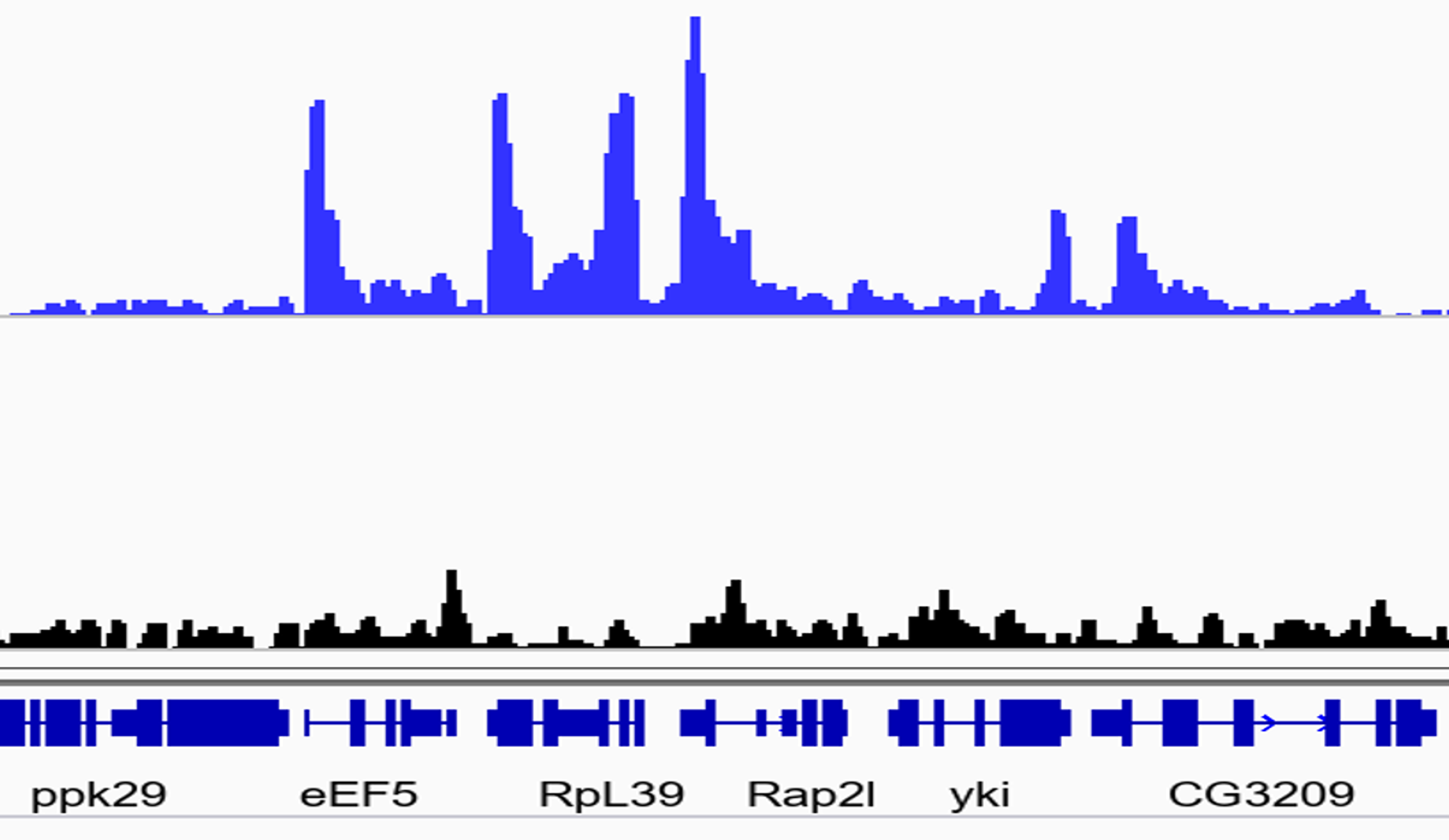
We recently published that Drosophila Sirt6 is a chromatin-associated, NAD+-dependent deacetylase, which removes acetyl residues from lysine 9 and lysine 56 on histone H3, much like mammalian Sirt6. Increasing Sirt6 levels dramatically alters gene expression, in particular by reducing expression of a subset of Myc-target genes, which are primarily involved in ribosome biogenesis. Emerging research suggests epigenetic modifier enzymes also play key roles in cellular function even when their primary catalytic activity is abolished. Our lab is currently investigating mechanisms of gene regulation by Sirt6. Topics of interest include: (i) identification of conserved Sirt6 target genes across multiple species, (ii) impact of histone deacetylation by Sirt6 on cis-gene expression, (iii) the role of Sirt6 catalytic activities in gene regulation , and (iv) identifying additional factors which mediate Sirt6 target gene specificity.